- This event has passed.
Prof. Cynthia Burrows ” Peering and Poking at RNA in Bacteria, Viruses and Vaccines”
September 23, 2022 @ 4:30 pm
Cynthia Burrows, Thatcher Presidential Chair and Distinguished Professor
Department of Chemistry, University of Utah, Salt Lake City
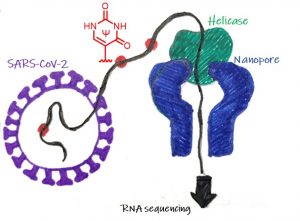
Nanopore-forming transmembrane proteins such as α-hemolysin provide the opportunity for electrophoretic capture and analysis of individual DNA or RNA strands in the protein cavity. Using an in-house device, we observed single-molecule dynamics of DNA mismatches, cytosine epigenetic modifications, and the unfolding of G-quadruplexes and i-motifs. Next, with the current availability of Oxford Nanopore Technology, we have been able to apply direct RNA sequencing for chemical modifications by inspecting the data for base-calling errors, ionic current and dwell-time differences, in comparison to the sequencing of control RNA without modifications. Application of these methods to viral RNA led us to identify conserved sites of pseudouridinylation of SARS-CoV-2 genomic RNA, and we could further sequence RNA synthesized by IVT in which m1Ψ replaces some of the U nucleotides with a view toward more effective mRNA vaccines. Finally, ribosomal RNA from natively grown E. coli cells that possess 36 chemical modifications of 17 different types were directly sequenced with the nanopore to identify unique base-calling errors, ionic current, and/or dwell-time signatures for 15 of the modification types, providing semi-quantitation of the modifications. The E. coli cells were then subjected to stressors followed by resequencing the rRNA to identify decreases in the levels of target base and ribose modifications. For example, the helicase dwell-time data permitted us to identify the single methyl group lost from N4-methyl-2`-O-methylcytidine in the 16S rRNA during thermal stress which triggers hibernation in E. coli. These results are a major step toward the Holy Grail of single-molecule sequencing for simultaneous identification and quantitation of all nucleotide chemical modifications in RNA.